提示词
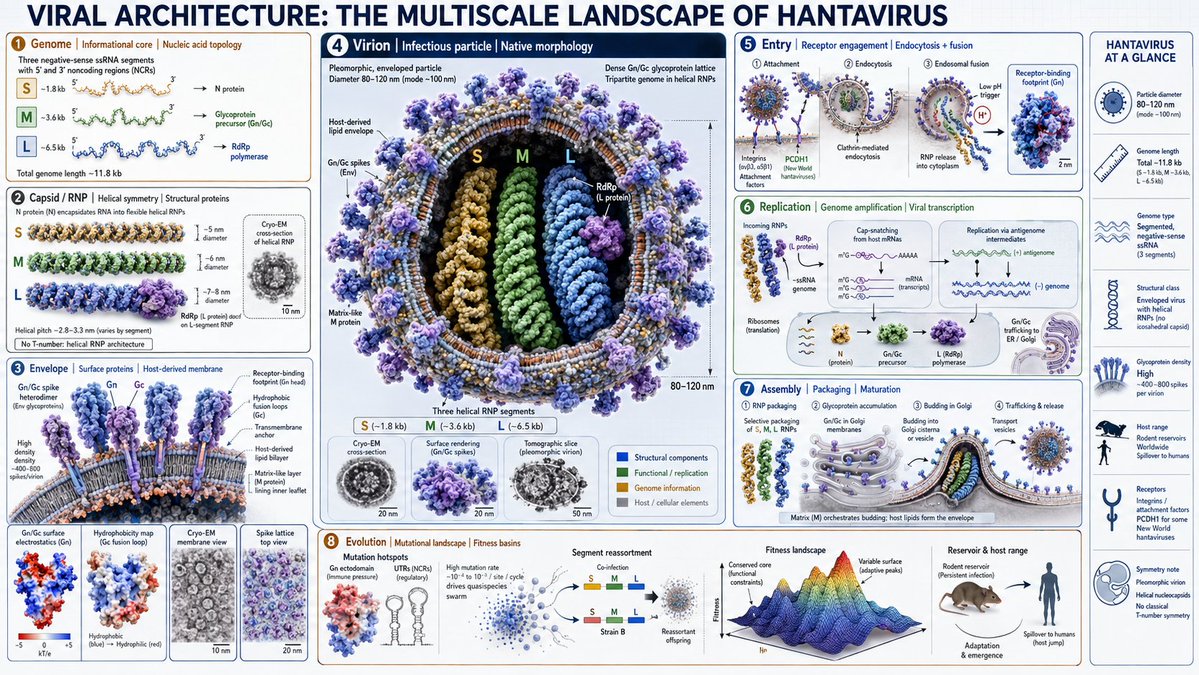
16:9, Do this for hantavirus => canvas: "Scientific Editorial Infographic" global_style: "Ultra-realistic 3D, cryo-EM inspired, nanoscopic macro-lighting, shallow DOF, virology aesthetic" title: "VIRAL ARCHITECTURE: THE MULTISCALE LANDSCAPE OF [$VIRUS]" layout_engine: type: "Hierarchical Viral Morphology + Assembly Pathway" connectors: "Energetic transitions, symmetry axes, genome–capsid interactions" frame_shape: "Icosahedral/Helical/Complex topology inferred from [$VIRUS]" flow_direction: "Genome → capsid → virion → host entry → replication cycle" frames: f1_genome: size: "xs" content: "Semantic inference of [$VIRUS] genome type (RNA/DNA, ss/ds, segmented/non-segmented)" ui_label: "Genome | Informational core | Nucleic acid topology" f2_capsid: size: "s" content: "Semantic inference of [$VIRUS] capsid architecture: icosahedral, helical, or complex symmetry" ui_label: "Capsid | Symmetry | Structural proteins" f3_envelope: size: "m" content: "Semantic inference of [$VIRUS] envelope state: lipid bilayer, glycoproteins, spikes, matrix layer" ui_label: "Envelope | Surface proteins | Host-derived membrane" f4_virion: size: "l" content: "Semantic inference of fully assembled [$VIRUS] virion: correct morphology, dimensions, and surface topology" ui_label: "Virion | Infectious particle | Native morphology" f5_entry: size: "m" content: "Semantic inference of [$VIRUS] host entry mechanism: receptor binding, fusion, endocytosis, or penetration" ui_label: "Entry | Receptor engagement | Membrane fusion or penetration" f6_replication: size: "m" content: "Semantic inference of [$VIRUS] replication strategy: Baltimore class logic, polymerase usage, replication compartments" ui_label: "Replication | Genome amplification | Viral factories" f7_assembly: size: "l" content: "Semantic inference of [$VIRUS] assembly pathway: capsid formation, genome packaging, budding or lysis" ui_label: "Assembly | Packaging | Maturation" f8_evolution: size: "xs" content: "Semantic inference of mutation hotspots, quasispecies dynamics, and structural constraints" ui_label: "Evolution | Mutational landscape | Fitness basins" side_panel: "Capsid triangulation numbers (T), helical pitch, envelope glycoprotein density, genome length, particle diameter, host range icons" halo_details: "Cryo-EM cross-sections, surface electrostatics, hydrophobicity maps, symmetry axes, receptor-binding footprints. Reject generic DNA helices, pill bottles, or non-viral artifacts."
中文版
相关推荐

A detailed scientific educational infographic about [Insert Creature, e.g., a Wo
![instructions> [SUBJECT]=Coronavirus. A hyper-realistic 3D zo](https://img.xuyi.dev/x-prompts/20260504/gdgtify-2051288232613351571-1.jpg)
instructions> [SUBJECT]=Coronavirus. A hyper-realistic 3D zoom-sequence infograp
![Render_Target = ( Architectural_Cutaway_3D_Model_Of_[DISH] *](https://pbs.twimg.com/media/HHZyI-gWoAAE1D6.jpg)
Render_Target = ( Architectural_Cutaway_3D_Model_Of_[DISH] * 1.5 ) + ( Ingredien

Create a high-end scientific infographic in a clean 2x2 grid layout featuring fo